Our mission at metaphacts has always been to ease the onboarding into the world of enterprise knowledge graphs. With our product metaphactory we provide an end-to-end platform to support that mission and enable our clients in unlocking the value of their data assets. Since we first published metaphactory in 2015, with every new release we have introduced new features and capabilities to enable rich end-user experiences in interacting with knowledge graphs.
Through our blog, we want to continuously share some of the recent developments, examples, best practices and make the power of knowledge graph technologies more accessible for you.
Just today we released metaphactory 3.6, so this is a great opportunity to start this blog with showing you some cool new additions to our product. With our most recent release, we have introduced a series of new components and enhancements that help provide a more intuitive user experience and user interaction. These new components cater to user needs across all platform target user groups: end users, developers focused on building end-user oriented applications, as well as knowledge graph experts.
Our Wikidata Demo System
To make the experience as concrete as possible, we provide live examples using the most prominent publicly available knowledge graph: Wikidata. For that purpose, we are hosting an instance of our metaphactory platform on top of Wikidata for you to experience. (If you have not heard about Wikidata yet, I recommend this short introduction, or this great article by Denny Vrandečić and Markus Krötzsch if you are interested in more detail and history.)
On the start page, you find a simple search box that allows you to search for any entity described in Wikidata. Perhaps you want to start by searching for your home town or anything of your interest. (Wikidata currently contains 90 million entities across many different domains - the coverage of Wikidata is what makes it a great resource for demonstrating the idea and value of a knowledge graph.)
The start page also contains entry points to more advanced use cases in various application domains as well as an example collection that shows the range of metaphactory components with the Wikidata Knowledge Graph.
Keyword Search featuring Type-based Disambiguation
In this post, we want to highlight one of the new search components, which aims to empower end users and deliver more flexibility in addressing hybrid information needs. The component supports simplified interaction with the data by allowing end users to combine keyword searches with type-based queries. Results can be visualized and explored flexibly and in context, with end users able to specify which properties should be visualized based on their relevance and the users' personal needs.
The structure and experience of the search interface, including the disambiguation and contextualization of search results, is driven by the underlying ontology of the knowledge graph, in particular its classes and their relationships. The Wikidata ontology is rather large and broad, but the search interface can be configured to focus on a specific part of the ontology that is of interest. In our example, we show a targeted search interface for the life sciences, with the following key classes and relationships that are of interest in use cases such as drug discovery:
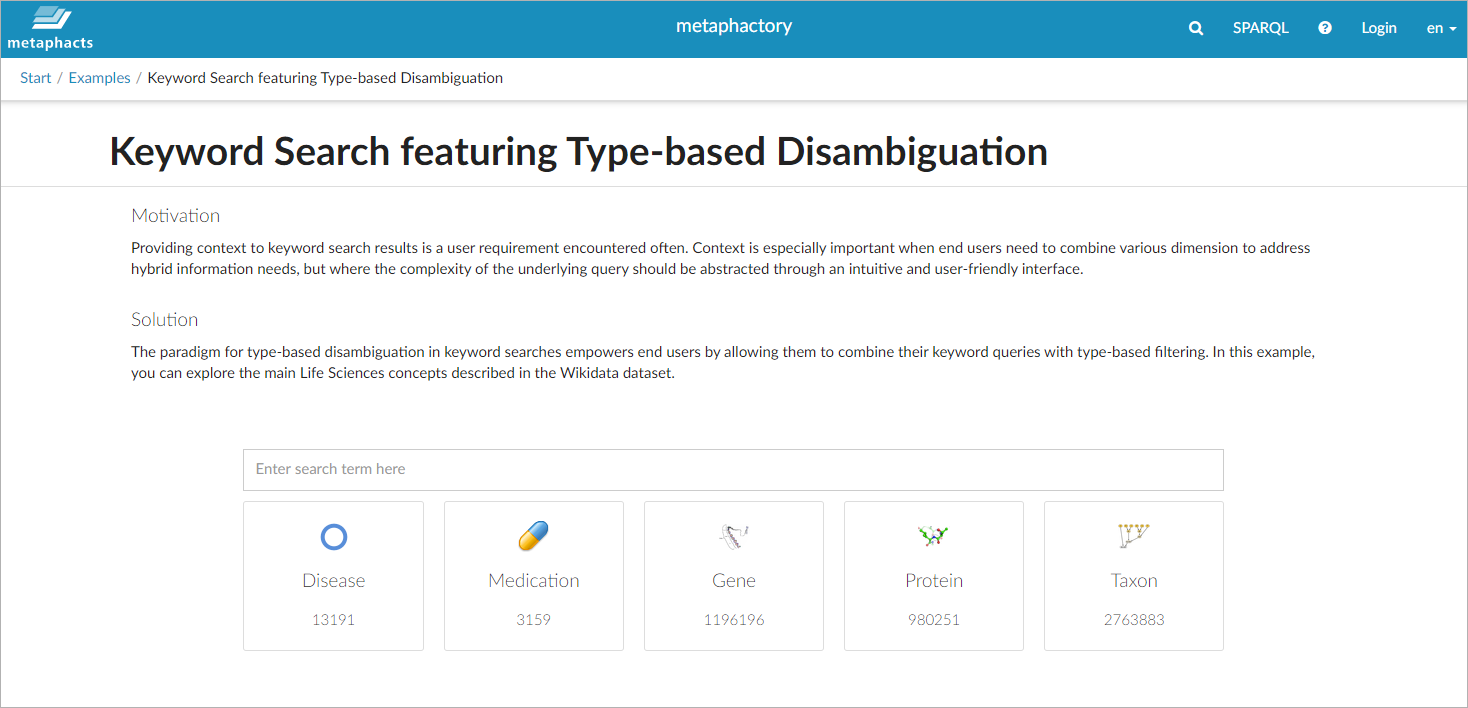
Specifically, we see the classes medication, disease, gene, protein, and taxon. The corresponding search interface is accessible on our Wikidata demo system »
It holds an input box for keywords and shows the main classes:
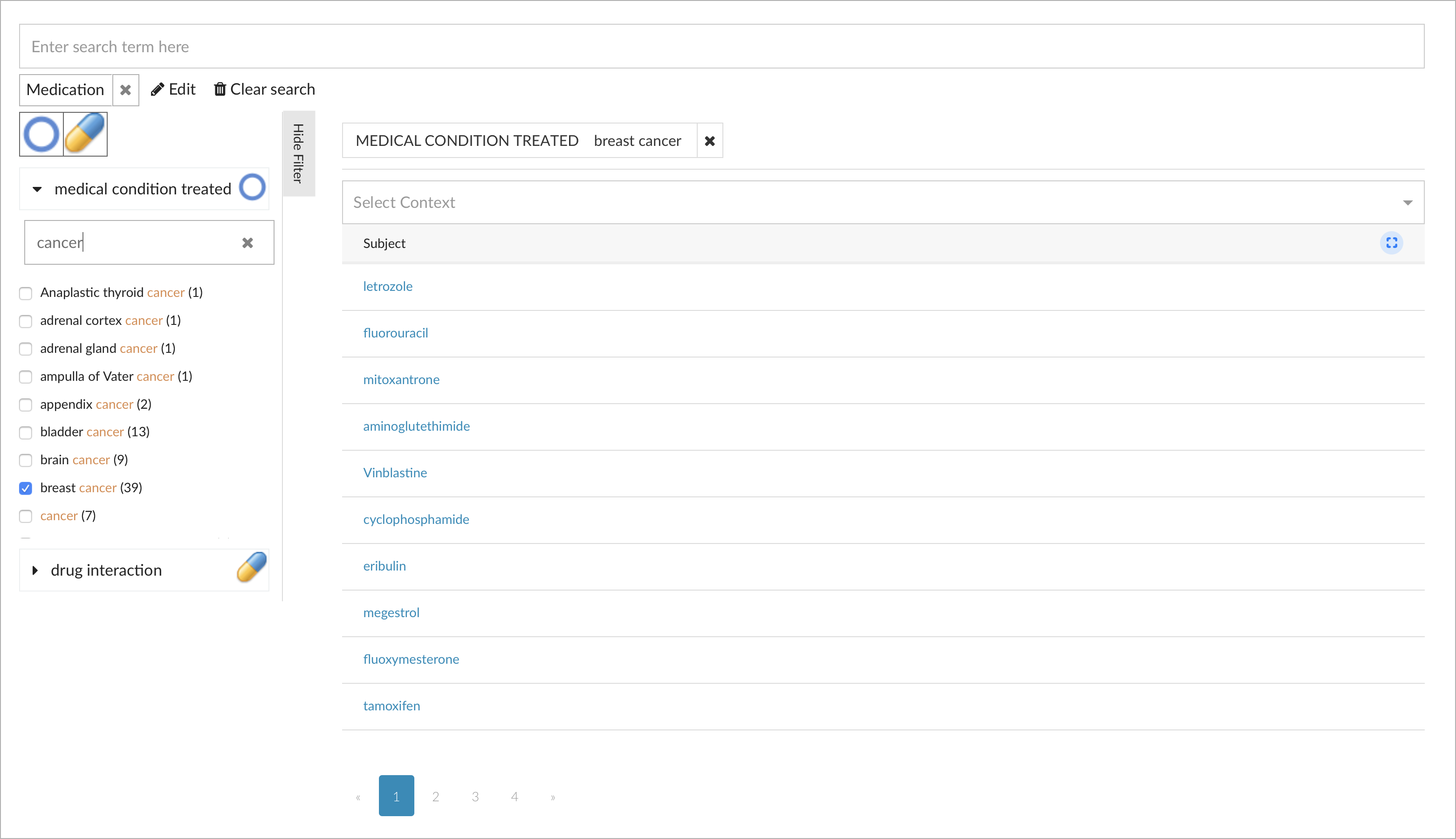
A search can be initiated by entering keywords, or simply by selecting a type to show all instances of that class, for example all medications. Facets, which are also computed dynamically based on the above model, allow to filter further, for example to search for medications to treat "breast cancer":
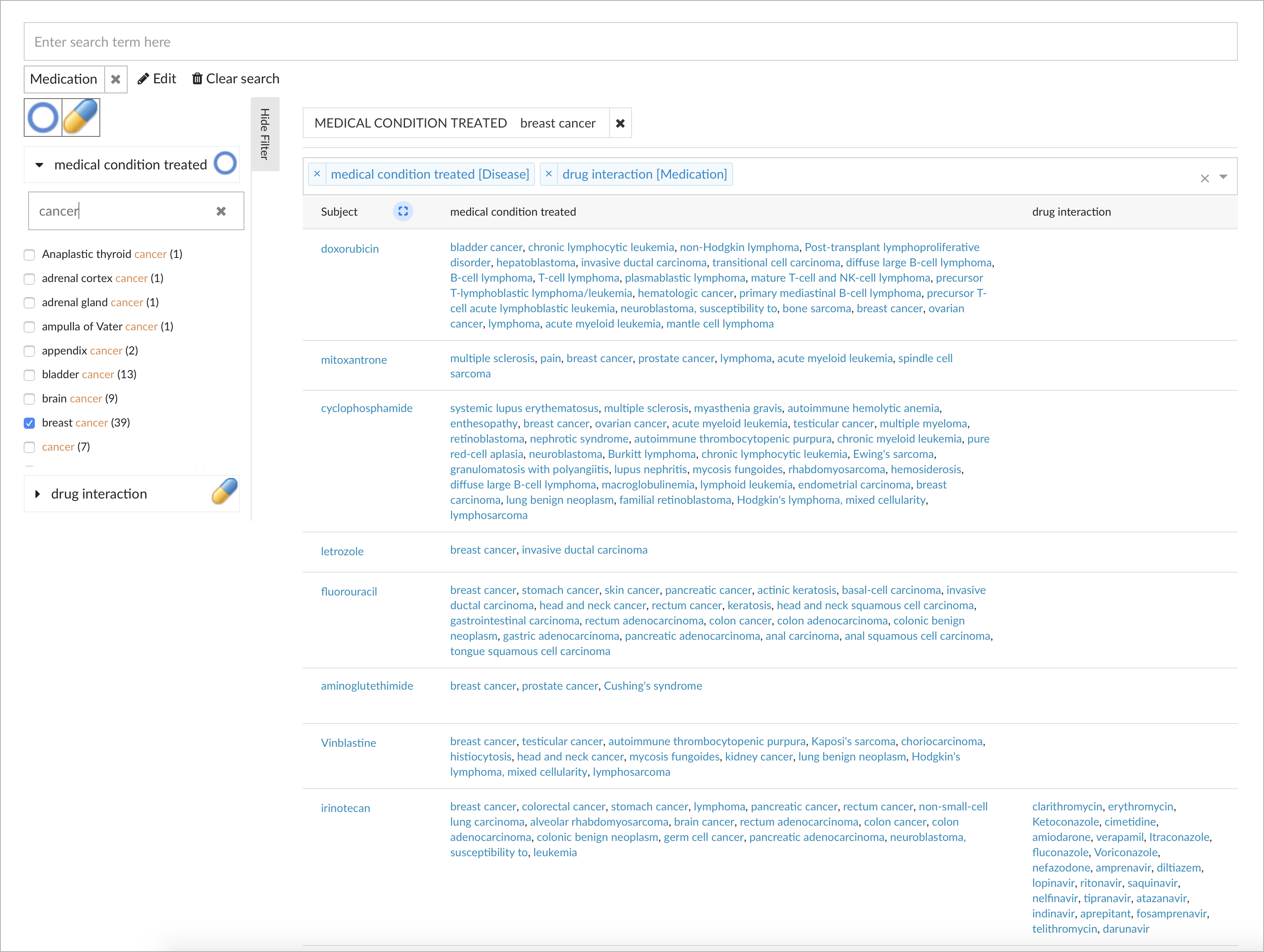
The result visualization in the table can easily be adjusted to show additional dimensions as defined in the model. For example, the result table showing the matching medications can be augmented with additional columns to show all medical conditions treated along with drug interactions, as defined by the relationships in the model:
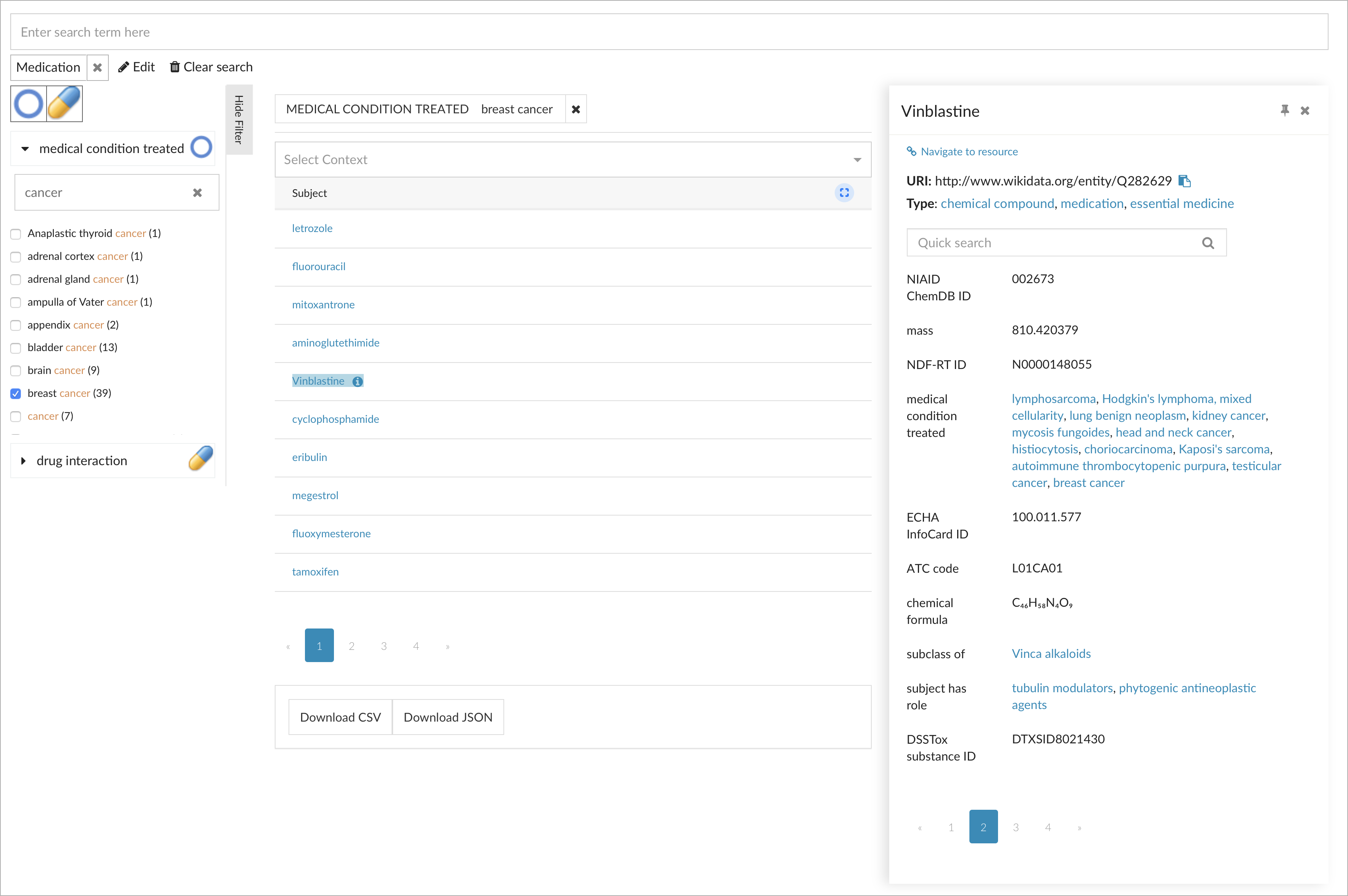
Much more detailed information is shown through the Knowledge Panel: The Knowledge Panel, just introduced as a generic component in our current metaphactory 3.6 release, allows to display a comprehensive structured summary of knowledge graph resources. As you hover over a result item, an icon appears to trigger the knowledge panel, here shown for the drug Vinblastine:
Without leaving the current context or navigating away to a separate page, you can easily get more details about a particular resource in place. Of course, the Knowledge Panel can be configured to show custom views depending on the type of resource.
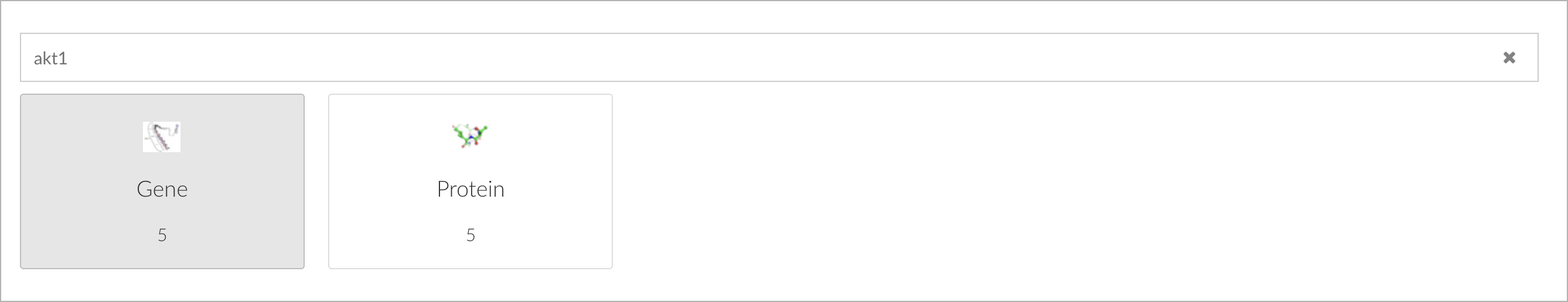
Another strong capability of the new search component is the ability to disambiguate. Search terms may mean different things in different contexts depending on information needs. For example, a user may be interested in a particular protein-coding gene in cancer research: AKT1. As you type a keyword into the search box, the search interface will indicate how many matches of each type have been found; in the case of AKT1 there are 5 genes and 5 proteins. You can disambiguate the search term by selecting a specific type, in our example we are looking for gene-related information:
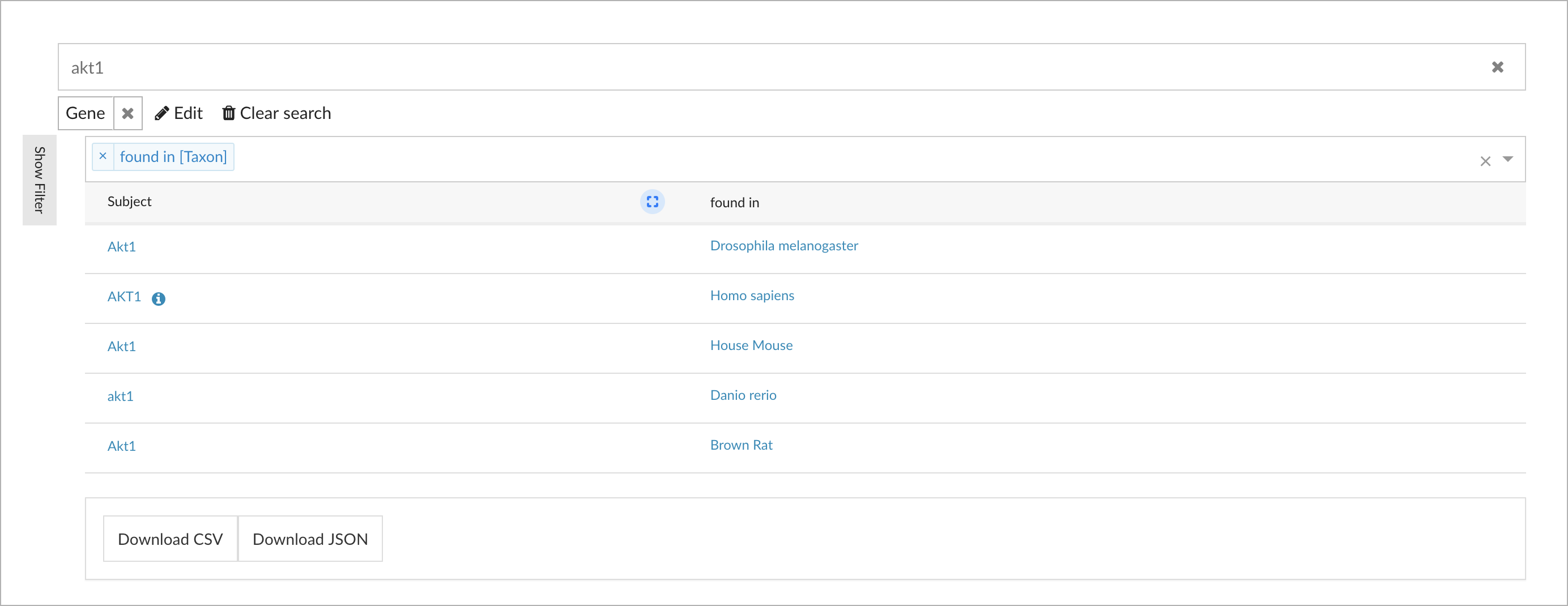
When the type has been selected, the corresponding results are shown. Even after specifying that we are interested in the gene AKT1, there are still five matches of genes with that exact name. Our model provides additional context that can be used for disambiguation:
You see the column 'found in' added as context in the result table, which shows the relationship with relevant taxa, such as 'Homo sapiens', and which easily allows to identify the correct protein-coding gene AKT1.
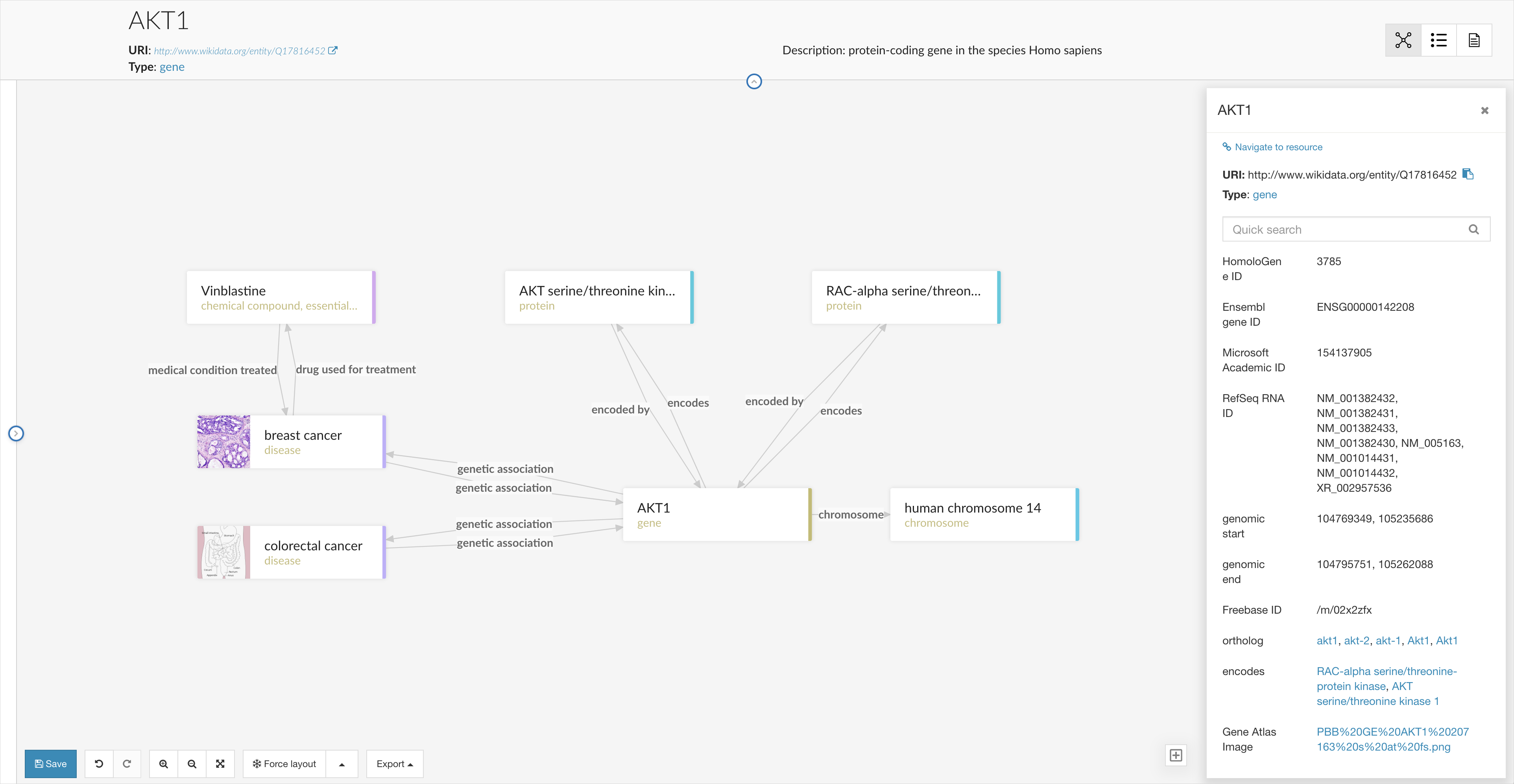
Of course, the disambiguated search result is only the starting point for much more detailed analysis supported with metaphactory - for example, as you navigate to a result page, easily explorable through our redesigned graph visualization and exploration component:
What you can do with this powerful component will be a great topic for a future blog post.
That's cool! How can I try it?
As mentioned already above, our live Wikidata demo system is accessible with many other examples here, so feel free to try the example described above or maybe start your search with diseases instead of medications. But there is more: You can even run the Wikidata demo system on your own! The system is available as an app for the metaphactory platform at https://github.com/metaphacts/wikidata
You can check it out to understand how it works, customize it, extend it, or use it as a blueprint to build your own app.
Of course, the keyword-type search component is just one of many search components in our semantic search framework. In a next blog post, we will feature a comparative description of the range of components in this search framework. And there is much more to come!
So stay tuned - follow our blog using the RSS feed and subscribe to our newsletter »